nami_toys#
Demonstrating making toy datasets using nami_toys. For full reproducibility, the same seed will have to be used,
import torch
import matplotlib.pyplot as plt
from nami_toys import (
Checkerboard,
GaussianMixture,
GaussianRing,
GaussianShell,
ParameterisedGaussian,
TwoMoons,
TwoSpirals,
Standardiser,
make_generator,
)
Seeded generator#
Before showing the different toys, its useful to quickly mention a utility for reproducibilty.
The make_generator(seed) function returns a seeded torch.Generator, or None if no seed (so it can be passed through to any toys generator kwarg)
# simple generation
ds_example = TwoMoons().generate(1000, generator=make_generator(42))
print(ds_example)
ToyDataset(n=1000, d=2, labels={0, 1}, meta={'noise': 0.1})
Gaussian mixture#
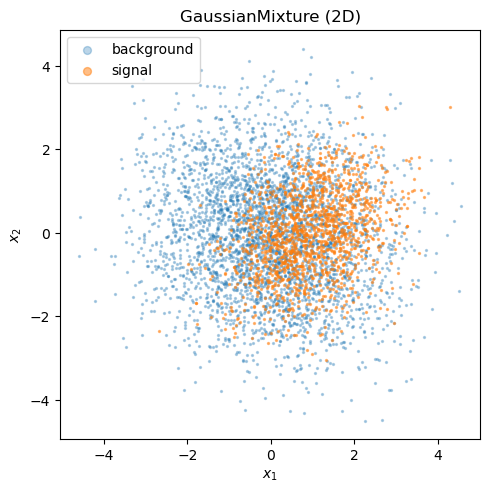
GaussianMixture draws signal and background events from two multivariate normals.
The total yield is Poisson-fluctuated and the signal/background split is binomial.
# use 2D defaults
gauss = GaussianMixture()
print(f"dimensionality: {gauss.d}")
print(f"signal mean: {gauss.sig_loc}")
print(f"background mean:{gauss.bkg_loc}")
dimensionality: 2
signal mean: tensor([1., 0.])
background mean:tensor([0., 0.])
ds = gauss.generate(5000, sig_frac=0.3)
print(f"events: {len(ds)}, shape: {ds.x.shape}, labels: {ds.y.unique().tolist()}")
print(f"meta: {ds.meta}")
events: 5072, shape: torch.Size([5072, 2]), labels: [0, 1]
meta: {'n_expected': 5000, 'sig_frac': 0.3}
sig_mask = ds.y == 1
bkg_mask = ds.y == 0
fig, ax = plt.subplots(figsize=(5, 5))
ax.scatter(*ds.x[bkg_mask].T, s=2, alpha=0.3, label="background")
ax.scatter(*ds.x[sig_mask].T, s=2, alpha=0.5, label="signal")
ax.set(xlabel="$x_1$", ylabel="$x_2$", title="GaussianMixture (2D)")
ax.legend(markerscale=4)
ax.set_aspect("equal")
plt.tight_layout()
Custom covariances#
You can pass custom location vectors and covariance matrices for arbitrary dimensionality to the GaussianMixture Dataset.
gauss_5d = GaussianMixture(
sig_loc=torch.tensor([2.0, 0.0, -1.0, 0.5, 0.0]),
sig_cov=torch.eye(5),
bkg_loc=torch.zeros(5),
bkg_cov=2.0 * torch.eye(5),
)
ds_5d = gauss_5d.generate(3000, sig_frac=0.5)
print(f"5-D dataset: {ds_5d.x.shape}")
5-D dataset: torch.Size([3032, 5])
Gaussian shell#
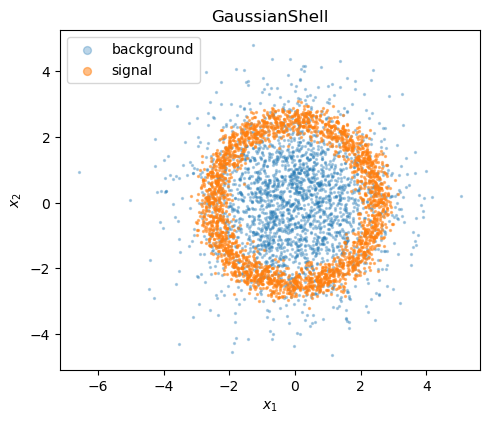
The GaussianShell dataset places signal events on a noisy ring and background from an isotropic 2-D Gaussian.
shell = GaussianShell(radius=2.5, width=0.25, bkg_scale=1.5)
ds_shell = shell.generate(5000, sig_frac=0.5)
print(f"events: {len(ds_shell)}, meta: {ds_shell.meta}")
events: 4999, meta: {'n_expected': 5000, 'sig_frac': 0.5, 'radius': 2.5, 'width': 0.25, 'bkg_scale': 1.5}
sig = ds_shell.y == 1
bkg = ds_shell.y == 0
fig, ax = plt.subplots(figsize=(5, 5))
ax.scatter(*ds_shell.x[bkg].T, s=2, alpha=0.3, label="background")
ax.scatter(*ds_shell.x[sig].T, s=2, alpha=0.5, label="signal")
ax.set(xlabel="$x_1$", ylabel="$x_2$", title="GaussianShell")
ax.legend(markerscale=4)
ax.set_aspect("equal")
plt.tight_layout()
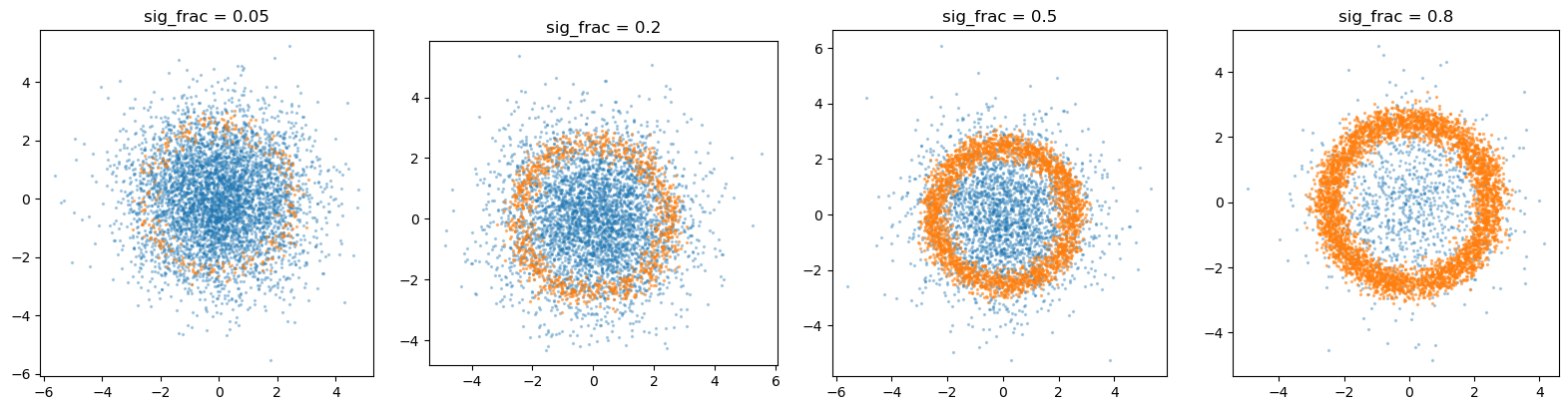
Varying the signal fraction#
The relative fraction of the two classes can be varied. A quick sweep over signal fractions is performed below to show how the dataset composition changes.
fracs = [0.05, 0.2, 0.5, 0.8]
fig, axes = plt.subplots(1, len(fracs), figsize=(4 * len(fracs), 4))
for ax, f in zip(axes, fracs):
d = shell.generate(5000, sig_frac=f)
s, b = d.y == 1, d.y == 0
ax.scatter(*d.x[b].T, s=2, alpha=0.3)
ax.scatter(*d.x[s].T, s=2, alpha=0.5)
ax.set_title(f"sig_frac = {f}")
ax.set_aspect("equal")
plt.tight_layout()
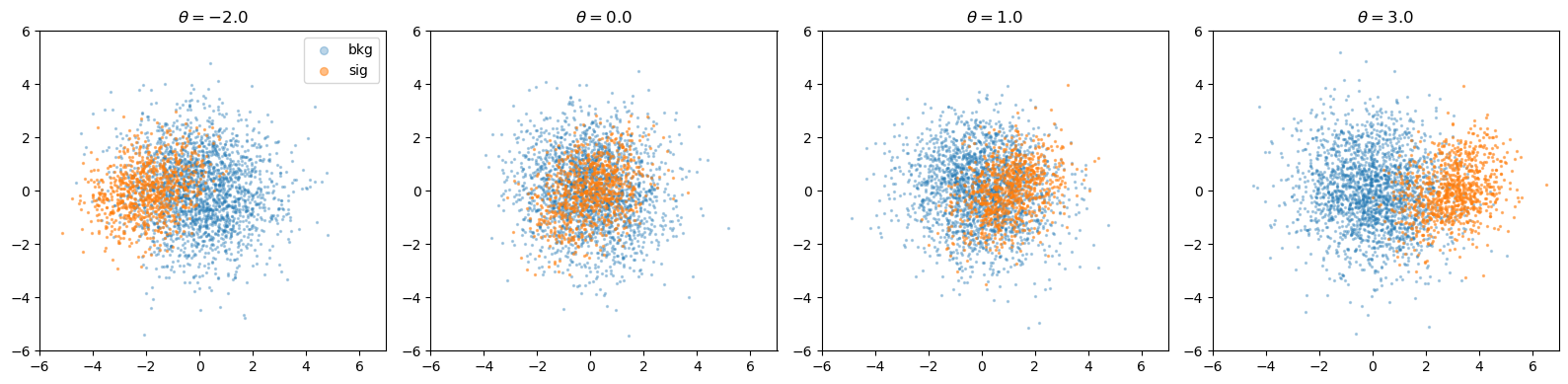
Parameterised Gaussian#
ParameterisedGaussian is a Gaussian mixture where the signal mean depends on a continuous parameter \(\theta\). The background stays fixed while the signal location shifts along a chosen dimension.
sim = ParameterisedGaussian(sig_frac=0.3)
print(f"dimensionality: {sim.d}")
print(f"param_dim: {sim.param_dim} (theta controls signal mean in this dimension)")
dimensionality: 2
param_dim: 0 (theta controls signal mean in this dimension)
thetas = [-2.0, 0.0, 1.0, 3.0]
fig, axes = plt.subplots(1, len(thetas), figsize=(4 * len(thetas), 4))
for ax, theta in zip(axes, thetas):
d = sim.generate(theta=theta, n_expected=3000)
s, b = d.y == 1, d.y == 0
ax.scatter(*d.x[b].T, s=2, alpha=0.3, label="bkg")
ax.scatter(*d.x[s].T, s=2, alpha=0.5, label="sig")
ax.set_title(rf"$\theta = {theta}$")
ax.set(xlim=(-6, 7), ylim=(-6, 6))
ax.set_aspect("equal")
axes[0].legend(markerscale=4)
plt.tight_layout()
The simulator also provides analytic log_prob and log_likelihood_ratio methods, handy for validating learned likelihoods.
x_test = torch.randn(5, 2)
print("log p(x | theta=1.0):", sim.log_prob(x_test, theta=1.0))
print("log-likelihood ratio:", sim.log_likelihood_ratio(x_test, theta=1.0))
log p(x | theta=1.0): tensor([-2.5743, -4.9410, -3.4000, -3.9989, -2.7842])
log-likelihood ratio: tensor([ 0.2119, -3.6674, -1.2144, -1.5441, 0.1194])
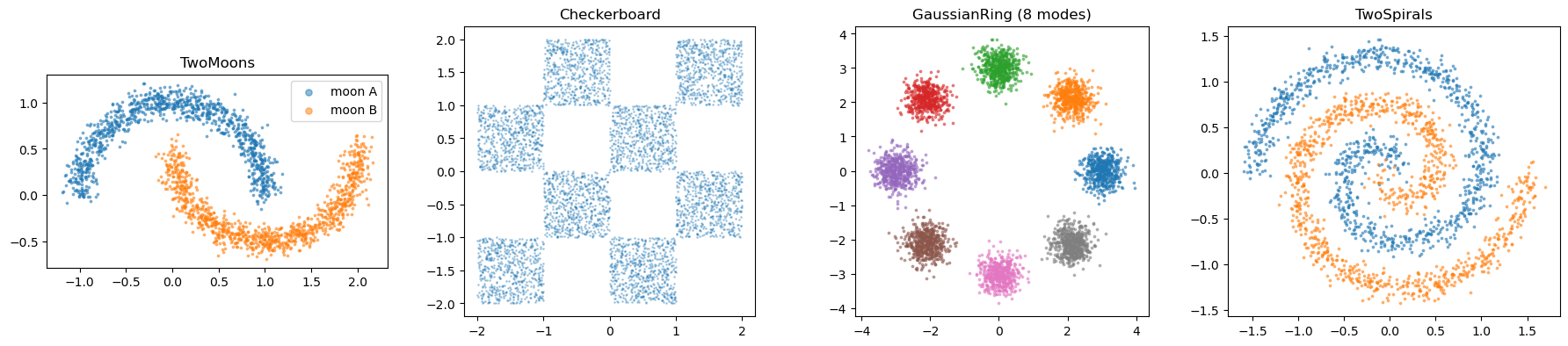
Density benchmarks#
The following toys generate samples from target densities.
fig, axes = plt.subplots(1, 4, figsize=(18, 4))
# two Moons
ds_moons = TwoMoons(noise=0.08).generate(2000)
for label, name in [(0, "moon A"), (1, "moon B")]:
m = ds_moons.y == label
axes[0].scatter(*ds_moons.x[m].T, s=3, alpha=0.5, label=name)
axes[0].set_title("TwoMoons")
axes[0].legend(markerscale=3)
axes[0].set_aspect("equal")
# checkerboard
ds_cb = Checkerboard(cells=4, bound=2.0).generate(5000)
axes[1].scatter(*ds_cb.x.T, s=1, alpha=0.3)
axes[1].set_title("Checkerboard")
axes[1].set_aspect("equal")
# ring of Gaussians
ds_ring = GaussianRing(n_modes=8, radius=3.0, std=0.3).generate(4000)
for mode in ds_ring.y.unique():
m = ds_ring.y == mode
axes[2].scatter(*ds_ring.x[m].T, s=3, alpha=0.5)
axes[2].set_title("GaussianRing (8 modes)")
axes[2].set_aspect("equal")
# two Spirals
ds_sp = TwoSpirals(noise=0.08, n_turns=1.5).generate(2000)
for label in [0, 1]:
m = ds_sp.y == label
axes[3].scatter(*ds_sp.x[m].T, s=3, alpha=0.5)
axes[3].set_title("TwoSpirals")
axes[3].set_aspect("equal")
plt.tight_layout()
ToyDataset utilities#
The ToyDataset container supports slicing via boolean masks and truncation.
# subset: keep only signal events
sig_only = ds_shell.subset(ds_shell.y == 1)
print(f"full: {len(ds_shell)}, signal only: {len(sig_only)}")
# limit: take first 200 events
small = ds_shell.limit(200)
print(f"limited: {len(small)}")
full: 4999, signal only: 2557
limited: 200
Feeding into a DataLoader#
Since everything is already a torch.Tensor, wrapping in a TensorDataset is trivial :)
from torch.utils.data import DataLoader, TensorDataset
loader = DataLoader(
TensorDataset(ds_shell.x, ds_shell.y),
batch_size=256,
shuffle=True,
)
x_batch, y_batch = next(iter(loader))
print(f"batch shapes: x={x_batch.shape}, y={y_batch.shape}")
batch shapes: x=torch.Size([256, 2]), y=torch.Size([256])
Standardiser#
## Standardiser
ds = GaussianShell().generate(5000, sig_frac=0.3)
scaler = Standardiser.fit(ds.x)
ds_z = scaler.transform_dataset(ds) # train on ds_z.x
print(f"standardised data: {ds_z}")
# samples_z = fm(None).sample((1000,)) # generate in standardised space
# samples = scaler.inverse(samples_z) # back to original scale
standardised data: ToyDataset(n=5137, d=2, labels={0, 1}, meta={'n_expected': 5000, 'sig_frac': 0.3, 'radius': 2.5, 'width': 0.25, 'bkg_scale': 1.5})